VT-SCC: Methodology
Introduction
VT-SCC employs an innovative dual-modal alignment architecture designed to dismantle the long-standing "species-specific isolation" (the "Siloed" paradigm) that restricts traditional agricultural counting. By facilitating a deep fusion of textual semantics and multi-scale visual representations, the system achieves precise identification and quantification of crop organs across diverse and complex morphologies.
Dependencies
Create and activate the conda environment:
conda create -n VT-SCC python=3.8 -y
conda activate VT-SCC
pip install torch==2.2.1 torchvision==0.17.1 torchaudio==2.2.1 --index-url https://download.pytorch.org/whl/cu121Required packages:
- timm==1.0.22
- transformers==4.39.1
- einops==0.8.1
- MultiScaleDeformableAttention==1.0
- pycocotools==2.0.8
- opencv-python==4.9.0.80
- pillow==11.3.0
- numpy==1.23.5
- pandas==2.3.1
- matplotlib==3.9.4
- tqdm==4.67.1
- PyYAML==6.0.1
- tensorboard==2.20.0
💾 Code and Data
1. Source Code: The source code can be downloaded from this link.
2. Dataset Availability: Only the test set of the M2-Crop (Multi-modal and Multi-crop Counting) dataset is publicly available for now. The training set will be released once the paper is officially accepted. You can download the dataset from this link.
3. Configuration: After downloading the dataset, please place it in your local directory and
modify the corresponding path in the ./config/datasets_all_test.json
file to specify your data path.
4. Pre-trained Weights: The pre-trained weights can be downloaded from this link.
🧠 Train
Train on M2-Crop training set:
sbatch train_all.sh🚀 Test
1. Test on M2-Crop test dataset
sbatch test_all.sh2. Test single image:
sbatch test_single_image.shResults
Consistency analysis confirms that the unified VT-SCC framework effectively handles morphological diversity, maintaining superior predictive stability across different species. By achieving R² scores of 0.9668 for wheat, 0.9724 for soybean, and 0.9906 for maize, the model outpaces specialized counterparts in single-crop tasks. Error distribution plots further show that our approach consistently maintains the lowest MAE and RMSE levels among all tested crops, with a unified MAE of 3.28. These results, notably the high precision in dense soybean scenarios (MAE 2.65), demonstrate the framework's success in leveraging scale-adaptive aggregation to resolve structural biases and align heterogeneous visual data.
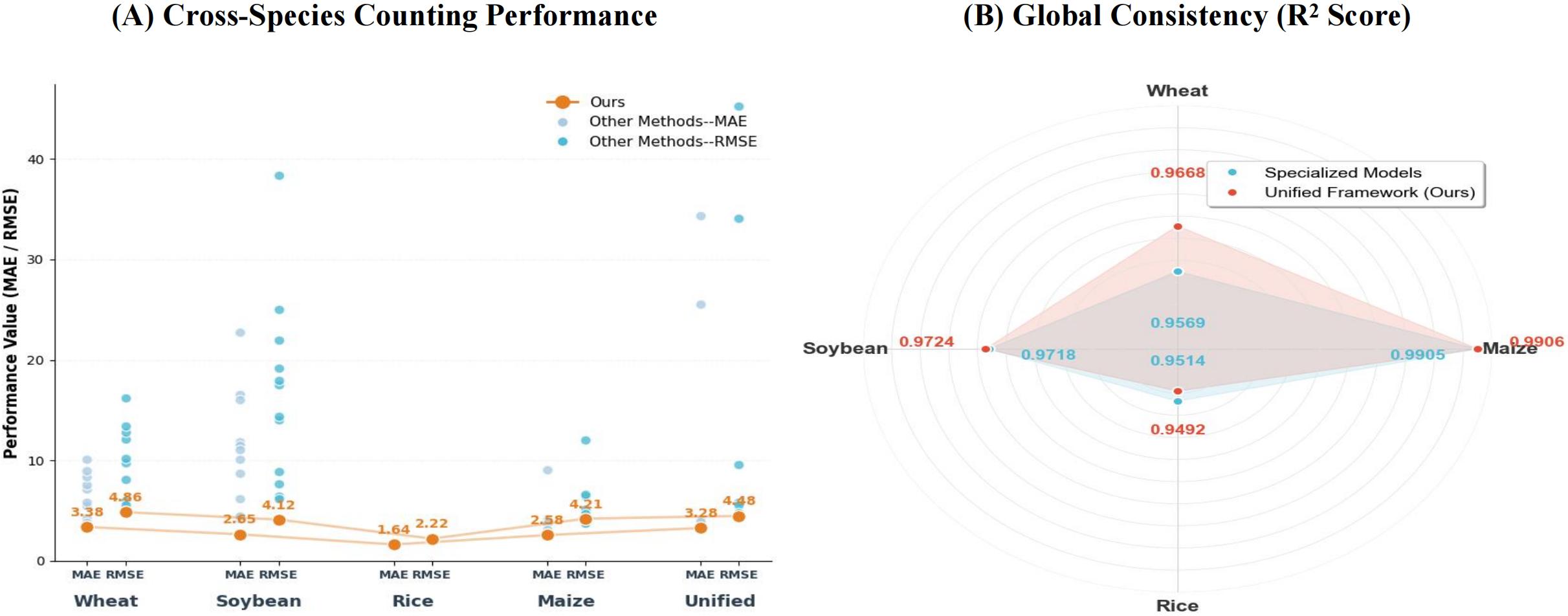
Figure 7: Evaluation of predictive consistency across diverse morphologies. (A) Scatter plots of MAE/RMSE trends; the consistently lower error profiles demonstrate effective multi-scale feature synergy. (B) Radar chart of R² scores; high stability across species verifies the capture of generalized crop invariants.
SAFA Feature Visualization
Feature activation visualizations provide direct evidence of SAFA's efficacy in coordinating multi-granularity representations. Observations indicate that, compared to the noisy and indiscriminate feature scatter of the raw backbone, SAFA-refined outputs achieve exceptional spatial precision by successfully shifting the perceptual focus from environmental clutter to target organs.
Figure 8: Comparative visualization of feature refinement via the SAFA module. (A) Raw input images; (B) Initial features from the backbone network, characterized by significant environmental noise and visual ambiguity; (C) SAFA-refined features, demonstrating high spatial precision by effectively shifting the focus to target organs. This transformation substantiates SAFA’s role in resolving scale-confused noise through multi-scale integration, establishing a high-quality semantic foundation.
SGCM Semantic Filtering
Visual analysis validates the efficacy of SGCM as a precise semantic filter. Results confirm that the model achieves high-fidelity focus on target organs while silencing background clutter under correct prompts, whereas mismatched instructions trigger a diffuse, low-confidence global search.
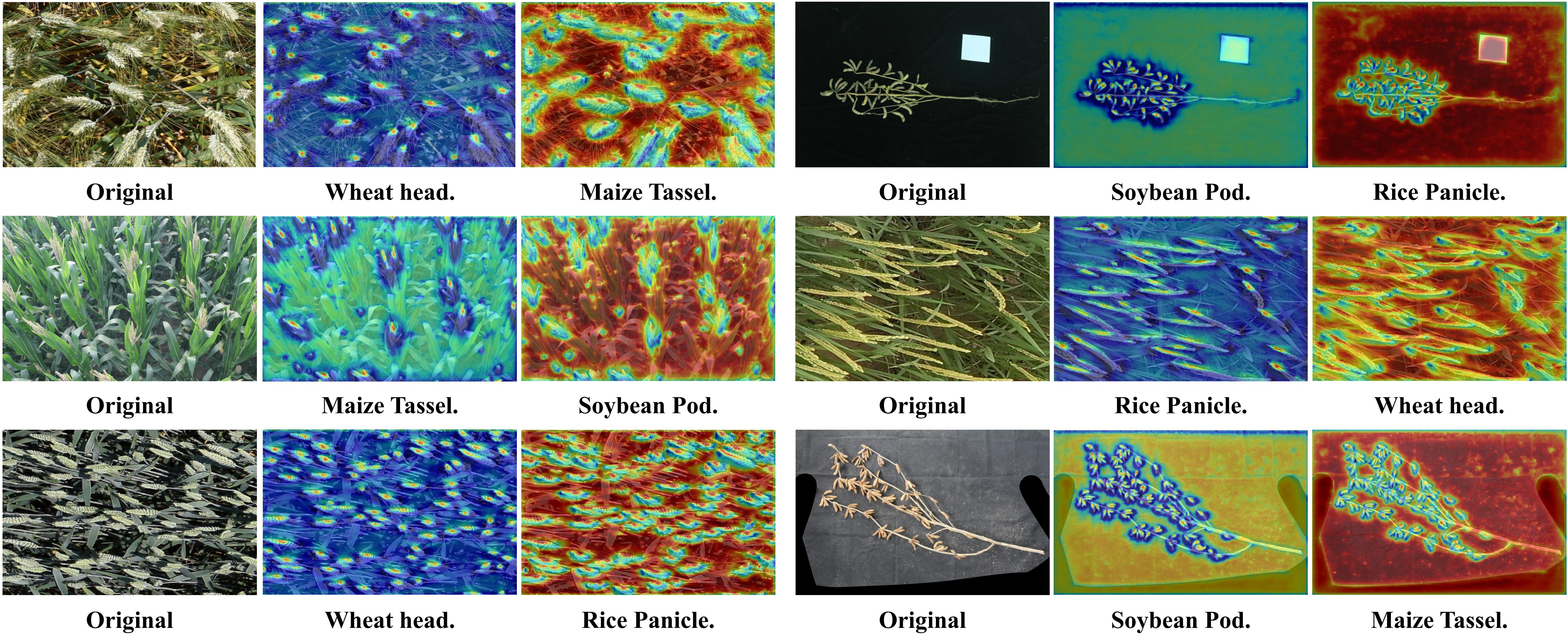

Figure 9: Qualitative assessment of the SGCM module under varying prompting conditions. The heatmaps visualize the spatial distribution of Modulation Intensity Δ driven by linguistic instructions. Under positive prompts, the module functions as a precise semantic filter, concentrating focus on targets; conversely, mismatched prompts trigger a diffuse search, validating the framework's strict sensitivity to task intent. This contrast demonstrates the efficacy of dynamic channel recalibration in isolating target signals from background clutter.
Regression Analysis
We utilized the coefficient of determination (R²) to evaluate the framework's predictive stability and alignment accuracy. The unified model attained an aggregate R² of 0.9755 across the entire test set. These findings confirm that the proposed paradigm ensures robust and globally consistent inference across diverse agricultural scenarios.
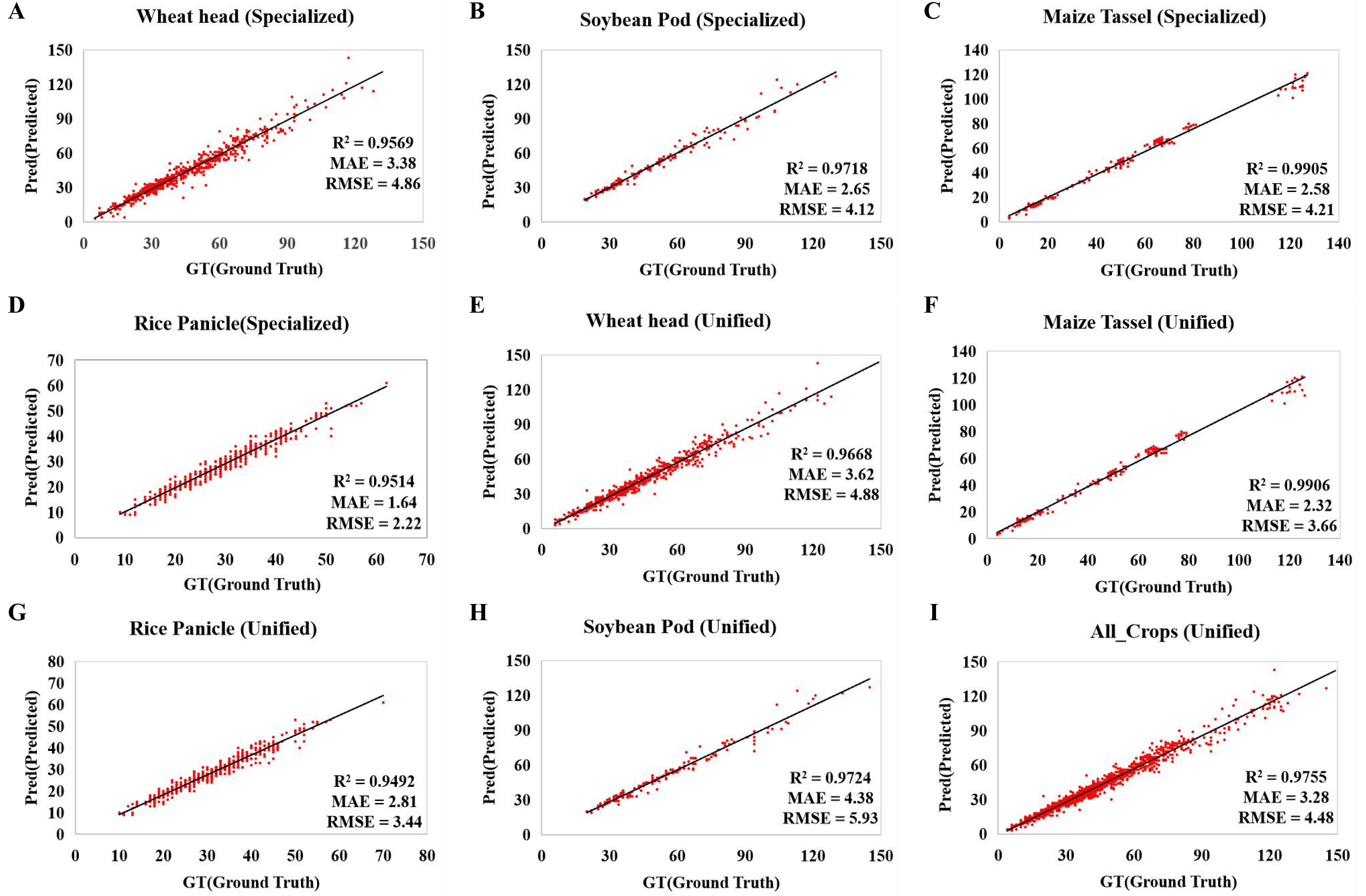
Figure 10: Regression analysis across counting paradigms. A-D: Specialized domain models. E-H: Performance of the unified framework on individual crops. I: Overall performance across the entire dataset. The consistently strong performance across all scenarios reinforces the framework's robustness.
